Viewing 3D Structures
ProtSpace integrates with AlphaFold to display 3D protein structures alongside your embedding visualization.
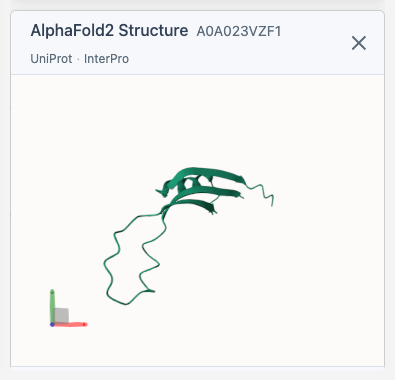
How It Works
When you select a protein with a UniProt accession:
- The structure viewer appears in the sidebar below the legend
- Links to AlphaFold Database, UniProt, and InterPro appear at the top - click them anytime
- The AlphaFold structure loads automatically via 3D-Beacons API
Supported Structures
Currently, ProtSpace supports AlphaFold structures only. PDB experimental structures are not yet integrated.
Confidence Coloring (pLDDT)
Structures are colored by predicted Local Distance Difference Test (pLDDT) confidence scores—the same scheme used on the AlphaFold Database. Regions in blue are high-confidence, yellow moderate, and red low-confidence. This helps you quickly spot which parts of the model are more reliable.
Viewer Controls
| Action | Effect |
|---|---|
| Left drag | Rotate the structure |
| Right drag | Pan the view |
| Scroll | Zoom in/out |
| Double-click | Reset the view |
When Structures Aren't Available
Not all proteins have AlphaFold structures. When no structure is found, the viewer displays:
"No 3D structure was found for <Protein ID>"
Next Steps
- Exporting Results - Save your findings
- FAQ - Common questions